Researchers at the University of Alberta are proposing a system that uses the genetics of cannabis plants to create a more accurate classification protocol for clinical research and production.
According to the recent study published in the journal PLOS ONE, the vernacular of sativa-versus-indica is outdated and inaccurate due to the constant misuse of botanical names, extensive cross-breeding, as well as labelling during unrecorded crossbreeding between genetically dissimilar plants.
Instead, the researchers suggest that cultivars be grouped in genetically similar clusters.
This is the first in a series of studies proposing a new cannabis classification system that analyzes the genome of a plant and genetic correlations among commercially-available cultivars.
“Although the cannabis industry is rapidly advancing after the relaxation of legal restrictions in North America, the increasing number of THC-dominant strains, CBD-dominant strains, and balanced strains only adds confusion to the currently poorly understood genetic background of the thousands of varieties already in existence,” reads the study.
(Note: While the researchers use “strain,” “cultivar” is the preferred term for plant identification.)
Previous classification studies have focused on chunks of DNA and not the whole genome. DNA is a molecule that functions as the biological “instructions” of a cell, to create proteins or other molecules that make up the body of a living organism. In this case, weed.
“In addition, CBD-dominant strains and balanced strains (THC = CBD), which have gained increasing attention due to CBD’s use as a therapeutic have been omitted in recent classification studies,” reads the study.
Read more: Teen cannabis use not associated with decline in cognitive skills, research shows
Read more: Terpenes imitate and enhance cannabinoid activity: study
The study extracted and tested the DNA of 23 cultivars after tagging them based on the three typical so-called chemotypes available in the market: THC-dominant, CBD-dominant and balanced.
A chemotype describes subspecies of plants with the same form and structure, but that produce different quantities of chemical components in their essential oils.
Researchers correlated the genetic information from the different strains using Discriminant Analysis of Principal Components, a method of statistical analysis designed to identify clusters of genetically related individuals.
The study showed the cultivars could be grouped into five chemotype-aligned clusters based on common genetics.
“These results showed that the genetic differences between modern cannabis strains were at a whole-genome level and not limited to THC or CBD production.”
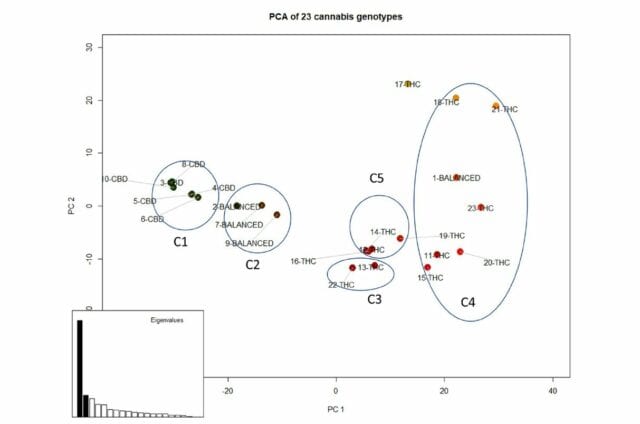
The 23 cultivars could be grouped into five chemotype-aligned clusters based on common genetics.
Five clusters with similar genetic properties were identified
The first cluster contained all six CBD-dominant cultivars. Another contained three out of four of the balanced cultivars.
The THC-dominant clusters were separated in three, with two clusters having two and three THC-dominant cultivars and the last cluster with the remaining 13 THC-dominant cultivars and one balanced.
CBD-dominant and the balanced cultivars were genetically closer to each other than THC-dominant ones.
“This may be attributed to how medical interest in breeding for non-psychoactive, CBD-elevated strains (CBD-dominant and balanced strains) has only recently been in vogue, resulting in an overlapping and less diverse gene pool for CBD-dominant and balanced strains compared to the longer breeding history for THC strains,” reads the study.
The study tested commercially available cultivars including Lemon Garlic OG, Royal Medic, Blue Hawaiian, Kandy Kush, Lemon Sorbet and Lemon Love.
Where possible, scientists reported ancestry as sativa-dominant or indica-dominant, which was taken from the licensed producer providing the weed or from Leafly’s online database.
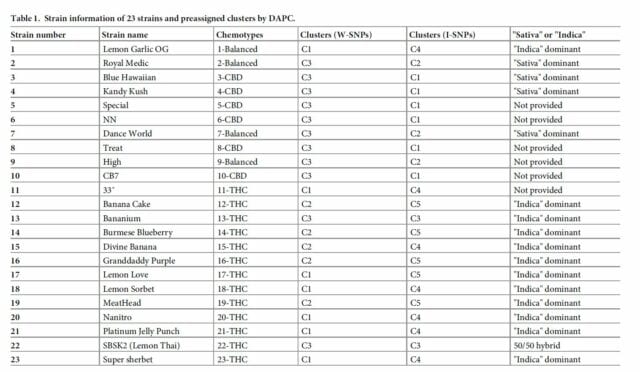
Cultivars that were tested for the study.
The study analyzed the whole DNA but focused on single nucleotide polymorphisms (SNPs) as the markers for determining differences. In genetics, these are described as a deviation in DNA structure seen as the change of position of one or more nucleotides, the basic building blocks of DNA.
These deviations in the DNA are useful for understanding the genetic basis of diversity among populations, and identifying the potential for disease in an organism. SNPs are like mutations but occur more often.
A different study found only a moderate correlation between reported sativa or indica ancestry and genetic structure.
Future research will focus on how SNPs are associated with the production of other cannabinoids, terpenes, flavonoids, as well as other compounds and characteristics.
Researchers from the U of A study encourage other scientists to share genetic information on cannabis to increase the available database and help build a more accurate system.
Follow Natalia Buendia Calvillo on Twitter
natalia@mugglehead.com